/*!
* @class AlignmentGraph
*
* @extends Graph
*
* @headerfile <seqan/graph_align.h>
*
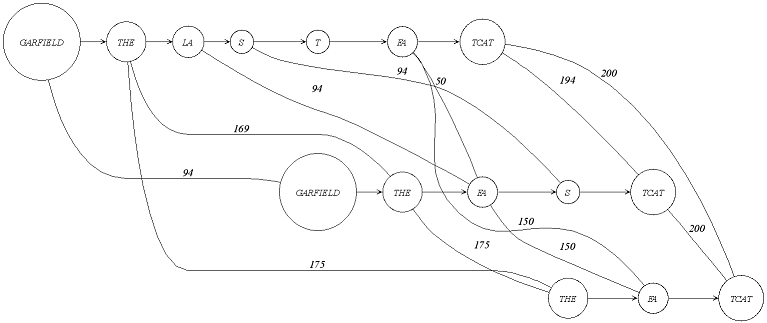
* @brief Alignment graph type.
*
* @signature template <typename TStringSet[, typename TCargo[, typename
* TSpec]]> class Graph<Alignment<TStringSet, TCargo, TSpec> >;
*
* @tparam TStringSet The type of the @link StringSet @endlink containing the
* sequence information. <b>Must be a @link
* DependentStringSet @endlink.</b>
* @tparam TCargo The cargo type that can be attached to edges. Default:
* <tt>unsigned</tt>.
* @tparam TSpec The specializing type. Default: <tt>Default</tt>. Use
* <tt>WithoutEdgeId</tt> here to omit edge ids. NB: if edges do
* not store ids then external property maps do not work.
*
* <img src="alignmentGraph.png" tite="An alignment graph with 3 sequences." />
*
* @section Example
*
* @include demos/graph_align/graph_align.cpp
*
* @code{.console}
* Score = -8
* Alignment matrix:
* 0 . :
* ATCGAATGCGGA
* | ||| |
* ACTCGTTGC--A
* @endcode
*
*
* @fn AlignmentGraph#getDistanceMatrix
*
* @headerfile <seqan/graph_msa.h>
*
* @brief Computes a pairwise distance matrix from an @link AlignmentGraph
* @endlink.
*
* @signature void getDistanceMtarix(graph, mat[, tag]);
* @signature void getDistanceMtarix(graph, mat[, kTup][, alphabet],
* KmerDistance);
*
* @param[in] graph An @link AlignmentGraph @endlink containing the sequences
* and possible alignment edges.
* @param[out] mat Pairwise distance matrix.
* @param[in] kTup For KMerDistance, the length of the k-mers.
* @param[in] alphabet For KMerDistance, the alphabet to use for k-mer counting
* (e.g. compressed alphabets).
* @param[in] tag See @link DistanceCalculationTags @endlink. Default:
* <tt>KMerDistance</tt>.
*
* @fn AlignmentGraph#assignStringSet
*
* @brief Assigns a new StringSet to an AlignmentGraph.
*
* @signature void assignStringSet(g, stringSet);
*
* @param[in,out] g An AlignmentGraph.
* @param[in] stringSet The @link StringSet @endlink to assign to.
*
* @see AlignmentGraph#getStringSet
* @see AlignmentGraph#stringSet
*
* @fn AlignmentGraph#getStringSet
*
* @brief Gets the StringSet of an AlignmentGraph.
*
* @signature THost getStringSet(g);
*
* @param[in,out] g An AlignmentGraph.
*
* @return THost A const reference to the StringSet's host.
*
* @see AlignmentGraph#assignStringSet
* @see AlignmentGraph#stringSet
*
* @fn AlignmentGraph#stringSet
*
* @brief Return reference to the StringSet of an AlignmentGraph.
*
* @signature THostRef stringSet(g);
*
* @param[in,out] g An AlignmentGraph.
*
* @return THost A reference to the StringSet's host.
*
* @see AlignmentGraph#assignStringSet
* @see AlignmentGraph#getStringSet
*
* @fn AlignmentGraph#label
*
* @brief Get the label associated with this vertex descriptor or the sequence
* that is associated with a fragment.
*
* @signature TInfix label(ag, v);
*
* @param[in] ag The AlignmentGraph to query.
* @param[in] v The vertex descriptor to query.
*
* @return TInfix An infix representing the sequence label.
*
* @fn AlignmentGraph#sequenceId
*
* @brief Gets the sequence id that is associated with this vertex descriptor or
* with a sequence of a fragment.
*
* @signature TId sequenceId(ag, v);
*
* @param[in] ag The AlignmentGraph.
* @param[in] v Vertex descriptor of the vertex to retrieve sequence id for.
*
* @return TId The sequence id.
*
* @fn AlignmentGraph#fragmentBegin
*
* @brief Gets the begin position for this vertex descriptor in the sequence.
*
* @signature TPos fragmentBegin(ag, v);
*
* @param[in] ag The AlignmentGraph to query.
* @param[in] v The vertex descriptor of the vertex.
*
* @return TPos The begin position.
*
* @fn AlignmentGraph#fragmentLength
*
* @brief Gets the length of the fragment for this vertex descriptor in the
* sequence.
*
* @signature TSize fragmentLength(ag, v);
*
* @param[in] ag The AlignmentGraph to query.
* @param[in] v The vertex descriptor of the vertex.
*
* @return TPos The fragment size.
*
* @fn AlignmentGraph#findVertex
*
* @brief Finds a vertx given a sequence id and a position.
*
* @signature TVertexDescriptor findVertex(ag, id, pos);
*
* @param[in] ag The AlignmentGraph to search in.
* @param[in] id The sequence id to search for.
* @param[in] pos The position to search for.
*
* @return TVertexDescriptor The vertex covering the given position on the
* specified sequence,
* <tt>getNil<TVertexDescriptor>()</tt> if none
* could be found.
*
* @fn AlignmentGraph#getProjectedPosition
*
* @brief Projects a position of one sequence taking part in a pairwise match
* onto the other sequence.
*
* @signature getProjectedPosition(ag, seqId, pos, seqId2, pos2);
*
* @param[in] ag The @link AlignmentGraph @endlink to query.
* @param[in] seqId The sequenceId.
* @param[in] pos The position to project from.
* @param[out] seqId2 The resulting id of the sequence that was projected onto.
* @param[out] pos2 The reuslting position of the sequence was was projected
* onto.
*
* @fn AlignmentGraph#getFirstCoveredPosition
*
* @brief Finds the first position in a sequence that is not assigned to a nil
* vertex.
*
* @signature TPos getFirstCoveredPosition(ag, id);
*
* @param[in] ag An @link AlignmentGraph @endlink.
* @param[in] id A sequence id.
*
* @return TPos A sequence position.
*
* @see AlignmentGraph#getLastCoveredPosition
*
* @fn AlignmentGraph#getLastCoveredPosition
*
* @brief Finds the last position in a sequence that is not assigned to a nil
* vertex.
*
* @signature TPos getLastCoveredPosition(ag, id);
*
* @param[in] ag An @link AlignmentGraph @endlink.
* @param[in] id A sequence id.
*
* @return TPos A sequence position.
*
* @see AlignmentGraph#getFirstCoveredPosition
*
* @fn AlignmentGraph#convertAlignment
*
* @brief Converts an alignment graph into an alignment matrix.
*
* @signature bool convertAlignment(g, component, order, compLength);
* @signature bool convertAlignment(g, matrix);
*
* @param[in] g AlignmentGraph to convert. An alignment graph.
* @param[out] component Vertex to component mapping.
* @param[out] order The order of the component graph when sorting
* topologically.
* @param[out] compLength Component sizes.
* @param[out] matrix A string that represents an alignment matrix.
*
* @return bool true iff the alignment graph is a valid alignment and false
* otherwise.
*
* @section Remarks
*
* The variant with <tt>component</tt> and <tt>order</tt> computes a topological
* sorting of connected components.
*/