Class
Gaps
Efficient storage of gaps for a sequence.
In the figure above, the source sequence has seven characters, the gapped sequence has four gaps and thus consists of eleven characters.
The gapped sequence is clipped to start at position 0 in the gapped sequence and to end at position 8 in the gapped sequence (the positions given as half-open intervals [begin, end) ).
Gaps wrap a SequenceConcept and allows to (1) insert gaps into the sequence and (2) select an infix of the gapped sequence (clipping).
The gaps are not inserted into the underlying sequence (source) but stored separately.
Using the clipping is optional and meant for selecting parts of the alignment as a part of the result of a local alignment algorithm.
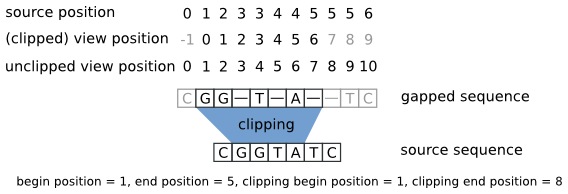 |
| Illustration of Gaps object and positions with clipping. |
The figure shows the three coordinate systems that are used with Gaps objects.
The source position is the position in the underlying sequence.
The unclipped view position is the position in the gapped sequence without gaps.
The view position is the position in the gapped sequence but including the clipping:
All (clipped) view positions have the clipping begin position subtracted from them.
Include Headers
seqan/align.h
Parameters
The type of the underlying sequence. Types: SequenceConcept | |
Specialization tag. |
Implements
Specializations
| Stores gaps anchors of the first characters behind gaps. | |
| Stores length of gap- and non-gapped runs in an array. |
Metafunctions
| Type for reading values. | |
| Determines whether a container stores its elements in sequential order. | |
| Type of iterator objects that are used to traverse the container. | |
| Type of an object that represents a position in a container. | |
| Reference type. | |
| Type of an object that is suitable to hold size information. | |
| Return underlying sequence of Gaps/Alignments. | |
| Type of the items in the container or behind an iterator. |
Member Functions
| Constructor |
Functions
| Assign the source of a Gaps object, copying data. | |
| Assigns value to item. (ContainerConcept) | |
| The last item in container. (ContainerConcept) | |
| Computes the best global pairwise alignment between two sequences given a non-empty seed chain. | |
| The begin of a container. (ContainerConcept) | |
| Begin position of object in host. (ContainerConcept) | |
| Clear clipping from Gaps object. | |
| Clear gaps and clipping from Gaps object. | |
| Return the begin position of the clipping in the unclipped gapped sequence. | |
| Return the end position of the clipping in the unclipped gapped sequence. | |
| The end of a container. (ContainerConcept) | |
| End position of object in host. (ContainerConcept) | |
| The first item in container. (ContainerConcept) | |
| Access to the value. (ContainerConcept) | |
| Computes the best global pairwise alignment. | |
| Insert one gap into a gapped sequence. | |
| Insert multiple gaps into a gapped sequence. | |
| Query whether a given clipped view position is a gap. | |
| The number of items/characters. (ContainerConcept) | |
| Computes the best pairwise local alignment using the Smith-Waterman algorithm. | |
| Assigns value to item. (ContainerConcept) | |
| Compute next suboptimal local alignment. | |
| Position of an iterator. (ContainerConcept) | |
| Remove one gap from a gapped sequence. | |
| Remove multiple gaps from a gapped sequence. | |
| Set the begin position of the clipped gapped sequence, given a source position. | |
| Sets the begin position of the clipping. | |
| Sets the end position of the clipping. | |
| Set the end position of the clipped gapped sequence, given a source position. | |
| Set the source of a Gaps object, do not copy if possible. | |
| Return underlying object. | |
| Compute split alignments. | |
| Transforms view to source position, if the view position is a gap, the original position of the next non-gap entry is returned. | |
| Transforms source to view position. | |
| Return length of the gapped sequence without clipping. |
Examples
The following example shows the construction of the gaps object from the image above together with some calls to toViewPosition and toSourcePosition .
These functions allow the transformation between the source position and the clipped view position.
File "gaps_example.cpp"
| 1 | |
| 2 | |
| 3 | |
| 4 | |
| 5 | |
| 6 | |
| 7 | |
| 8 | |
| 9 | |
| 10 | |
| 11 | |
| 12 | |
| 13 | |
| 14 | |
| 15 | |
| 16 | |
| 17 | |
| 18 | |
| 19 | |
| 20 | |
| 21 | |
| 22 | |
| 23 | |
| 24 | |
| 25 | |
| 26 | |
| 27 | |
| 28 | |
| 29 | |
| 30 | |
| 31 | |
| 32 |
This yields the following output:
toSourcePosition(gaps, 0) == 1
toSourcePosition(gaps, 4) == 4
toViewPosition(gaps, 0) == -1
toViewPosition(gaps, 5) == 9
SeqAn - Sequence Analysis Library - www.seqan.de
